DURHAM,
When things go awry -- the other 5 percent of the time -- it can affect development later in life, resulting in diseases like muscular dystrophy and related disorders in humans.
Scientists in Duke University’s School of Medicine have come one step closer to understanding how these crucial early stages of life are executed so perfectly most of the time by watching development of the fruit fly embryo.
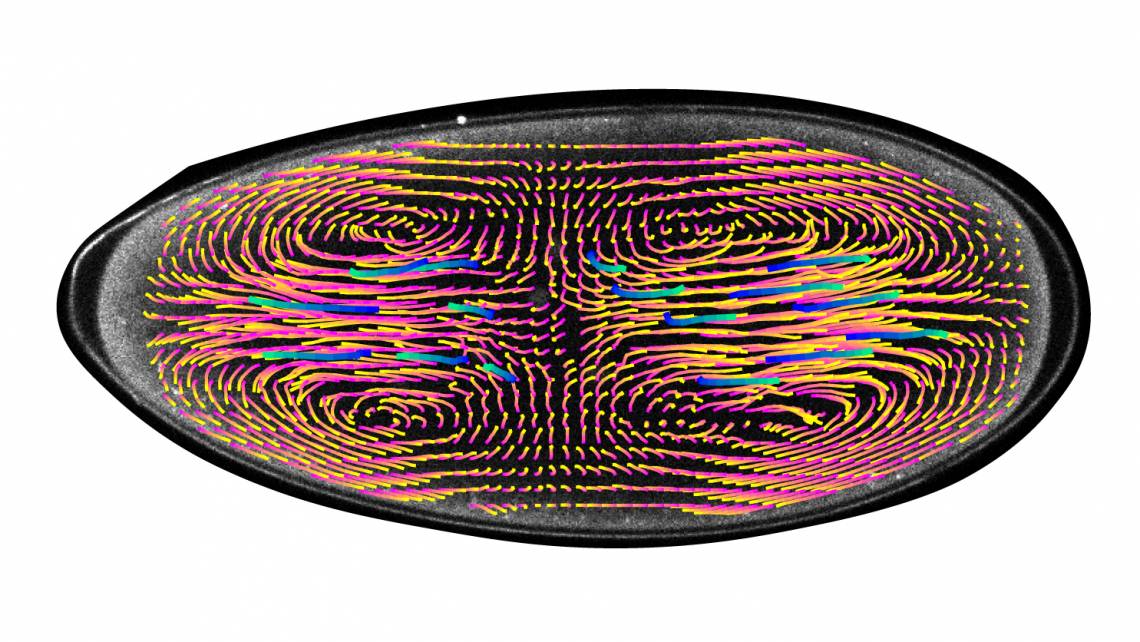
Unlike a human embryo, where a single cell multiplies through repeated cell divisions, the early embryo of a fruit fly starts as a football-shaped egg containing a single nucleus that divides into thousands of nuclei, all within the same cell.
Stefano Di Talia, PhD, an assistant professor in cell biology, and graduate student Victoria Deneke observed that oscillations of biochemical activity begin around the cell nuclei. These oscillations create currents that spread the nuclei into a specific formation that keeps them synchronized and allows them to undergo the same number of divisions. Previously, it was not known where these oscillations occurred and whether they had a role in controlling movements inside the embryo. The finding was published in the journal Cell on May 2.
“What really makes this paper special was our ability to put together all of the ideas that have been out there and create a model to solve the problem,” said Di Talia. “Using the sophisticated computational model we developed, we can determine exactly what the biochemistry is, and how that interfaces with cell mechanics.”
Using the computational model, which was developed with the expertise of co-author Alberto Puliafito at the University of Turin, the scientists were able to determine that if the process of nuclear positioning doesn’t happen properly, it will result in cell division falling out of synch and genetic defects.
“It does it so precisely because there is evolutionary pressure for this process to be controlled so that development can proceed accurately,” said Di Talia.
Even though fruit fly embryos develop differently than human embryos, the scientists said that the study can still shed light on human development and regeneration studies.
“Broadly speaking, there are a lot of commonalities between all animals,” said Deneke, first author on the paper. “We all have to expand our cell number first and then specify them into different cell types. And how that process is coordinated is essentially what we’re trying to learn. Our study reveals general principles that can be applied to more complex tissues.”
This research was supported by a Schlumberger Faculty for the Future Fellowship, an HHMI International Student Research Fellowship, a Boehringer Ingelheim Fonds Travel Grant, a Fondazione Umberto Veronesi postdoctoral fellowship 2017 and 2018, Early (P2ELP3_172293) and Advanced (P300PA_177838) Postdoc Mobility fellowships from the Swiss National Science Foundation, an Associazione Italiana per la Ricerca sul Cancro (AIRC AIG 18675), Fondazione Piemontese per la Ricerca sul Cancro, Ministero della Salute (FPRC 5x1000 2014 to L.P.), and NIH (R01-GM122936 to S.D.T.)
CITATION: “Self-Organized Nuclear Positioning Synchronizes the Cell Cycle in Drosophila Embryos,” Victoria Deneke, Alberto Puliafito, Daniel Krueger, Avaneesh Narla, Alessandro De Simone, Luca Primo, Massimo Vergassola, Stefano De Renzis, Stefano Di Talia. Cell, May 2, 2019. DOI: 10.1016/j.cell.2019.03.007. Online: https://www.cell.com/cell/fulltext/S0092-8674(19)30269-7